A classification model based on Agraph convolution, a classification model used to identify anti-cancer metabolites from the Vietnamese traditional herbal database

1. Article overview
For thousands of years, Vietnam has been a rich and diverse source of herbal medicine, playing various purposes in drug development to solve health problems such as cancer. According to the principles of chemoinformatics, compounds with similar structures are likely to have similar biological activities. This study uses molecular graph convolution and machine learning architecture to extract features from small molecules as undirected graphs to predict the structure of Vietnamese herbs based on metabolites. The anti-cancer ability. In addition to molecular map convolution, extended connection fingerprints, a traditional saturator using molecular details (ECFP), for performance comparison. Finally, we successfully constructed a neural network based on graph convolution with high prediction accuracy, indicating that the model is reliable in detecting anti-cancer activity.
Two, graphic guide
For thousands of years, Vietnam has been a rich and diverse source of herbal medicine, playing various purposes in drug development to solve health problems such as cancer. According to the principles of chemoinformatics, compounds with similar structures are likely to have similar biological activities. This study uses molecular graph convolution and machine learning architecture to extract features from small molecules as undirected graphs to predict the structure of Vietnamese herbs based on metabolites. The anti-cancer ability. In addition to molecular map convolution, extended connection fingerprints, a traditional saturator using molecular details (ECFP), for performance comparison. Finally, we successfully constructed a neural network based on graph convolution with high prediction accuracy, indicating that the model is reliable in detecting anti-cancer activity.
Two, graphic guide
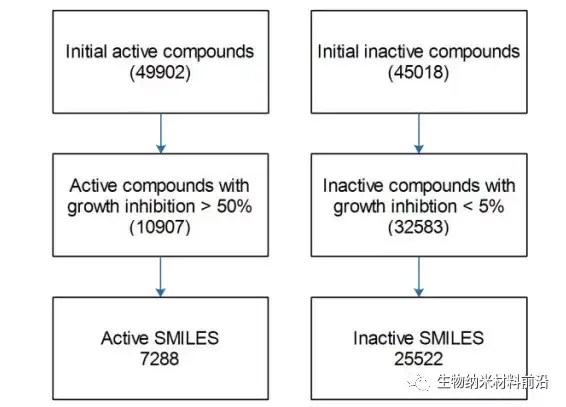
Figure 1. Building a data set for training and predicting anti-cancer activity from the NCI-60 data set.
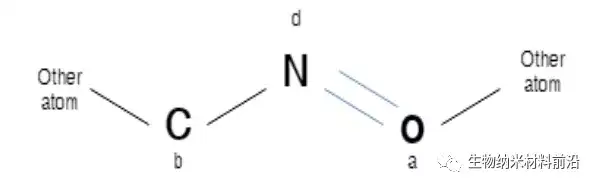
Figure 2. Explanation of some molecules at iteration 0. N represents a nitrogen atom, which is considered to be a detection atom or a core atom, and its identifier is d. C and O represent carbon, respectively, the oxygen atom with the identifier b and the oxygen atom with the identifier a. A row represents a single bond, and a double line represents a double bond.
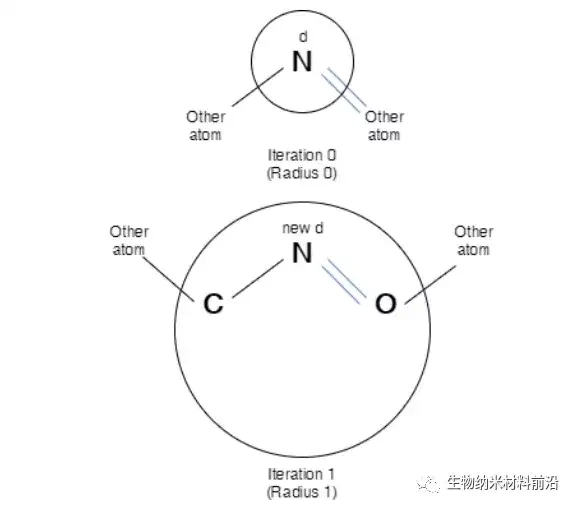
Figure 3. Description of the identifier d of N atoms in iteration 0 and iteration 1. After each iteration, the sub-structure representation becomes richer in terms of chemical structure insight.
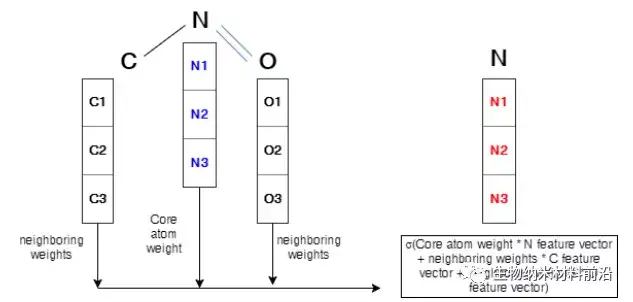
Figure 4. Operation description in the graph convolutional layer. The arrays of C1, C2, and C3 represent the feature vector of the C atom. The same symbols are used for N atoms and O atoms. The core atom, nitrogen, is marked in blue. In order to perform convolution to form a new feature of the core atom related to the 3 adjacent atoms, this sum will be carried out and wrapped by an activation function. The new feature vector is marked in red. Similarly, the same process will be applied to all nodes in the molecular structure, which will generate a brand new feature vector for each node.
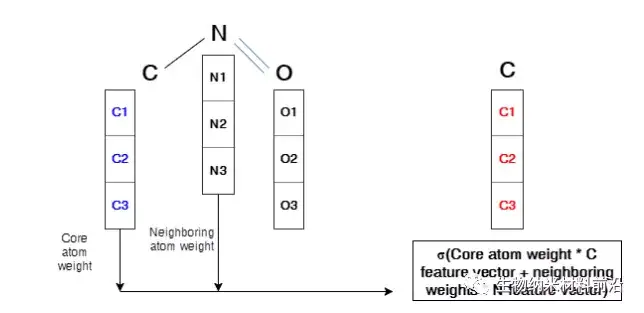
Figure 5. Illustration of the graph convolutional layer on the core atom of two adjacent atoms.
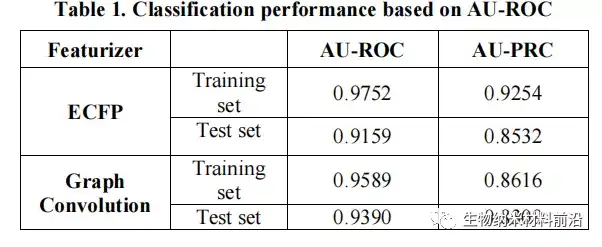
Figure 6. Based on the classification performance of Lazuli-Rosneft.
3. Full text summary
We successfully constructed a predictive model for anti-cancer activity. Despite the small size of the data set, the results are promising. In addition, we successfully grabbed the smile data from the NCI-60 database and filtered it to obtain the final cleaned data set, thus achieving good performance. In the future, the model will be constructed as an independent version as a tool for mining medicinal plants in Vietnam. Based on this research, DeepChem is not only capable of discovering anti-cancer drug candidates, but also can be used to map out potential drug candidates for other diseases.
Article link:
http://n.ustb.edu.cn/https/77726476706e69737468656265737421f4fb0f9d243d265f6c0f/doi/10.1145/3184066.3184090
This information is sourced from the Internet for academic exchanges only. If there is any infringement, please contact us to delete it immediately.
3. Full text summary
We successfully constructed a predictive model for anti-cancer activity. Despite the small size of the data set, the results are promising. In addition, we successfully grabbed the smile data from the NCI-60 database and filtered it to obtain the final cleaned data set, thus achieving good performance. In the future, the model will be constructed as an independent version as a tool for mining medicinal plants in Vietnam. Based on this research, DeepChem is not only capable of discovering anti-cancer drug candidates, but also can be used to map out potential drug candidates for other diseases.
Article link:
http://n.ustb.edu.cn/https/77726476706e69737468656265737421f4fb0f9d243d265f6c0f/doi/10.1145/3184066.3184090
This information is sourced from the Internet for academic exchanges only. If there is any infringement, please contact us to delete it immediately.
+86-18915413828(WhatsApp&WeChat)
Previous: Liu Peifeng Adv Sci: N


